 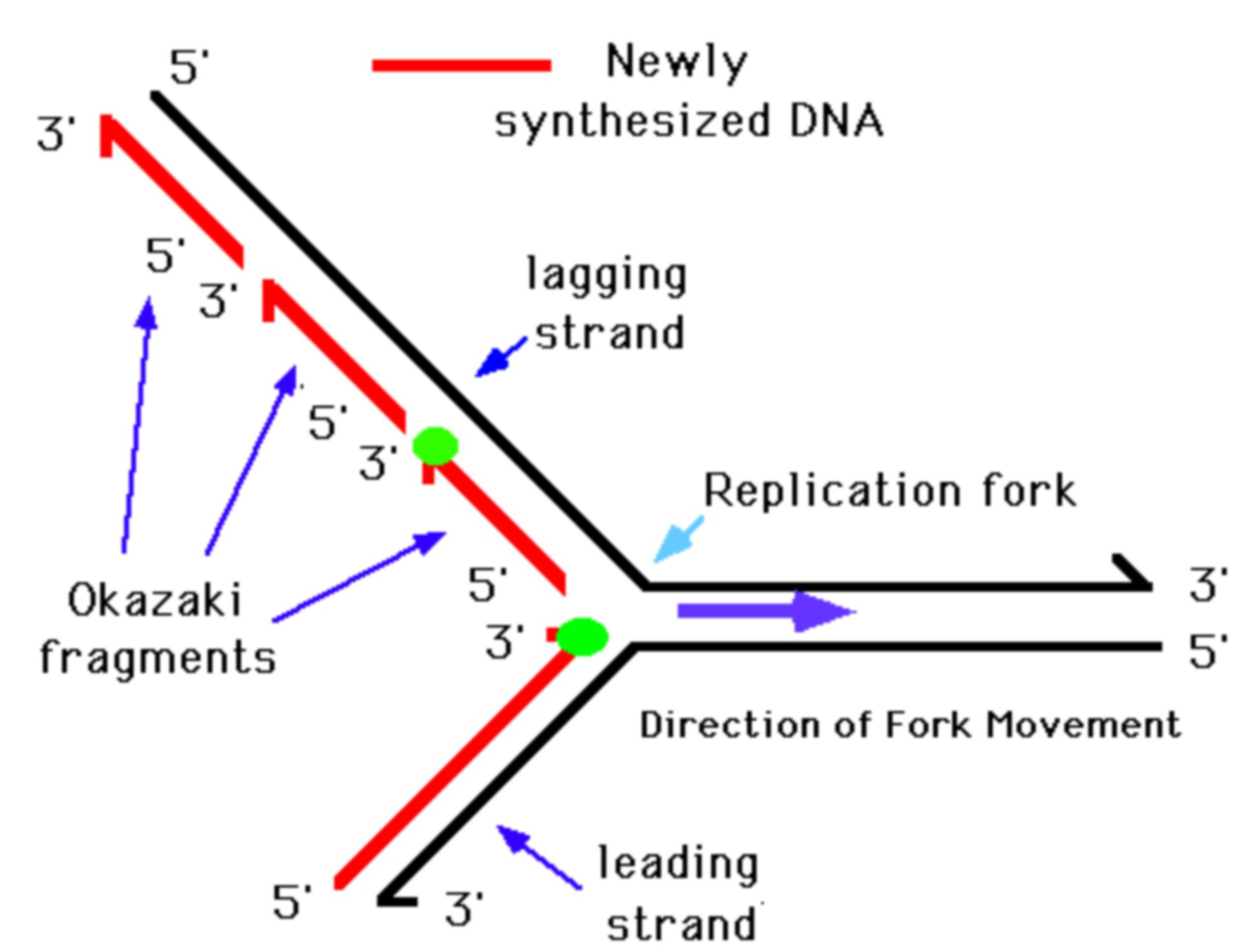
Unsuccessful processing of Okazaki fragments leads to the accumulation of DNA breaks which are associated with many human diseases including cancer and neurodegenerative disorders. The short DNA fragments are synthesized in the direction opposite to the replication fork processing. Cras mattis consectetur purus sit amet fermentum. Pellentesque ornare sem lacinia quam venenatis vestibulum. The Okazaki fragments definition are fragments of DNA produced during DNA replication. Okazaki fragments are the DNA fragments that result from the discontinuous replication of the lagging strand. 
Okazaki fragments are pieces of DNA that are transient components of lagging strand DNA synthesis at the replication fork. Nonetheless, the polA1 strain was viable under a standard culture condition. The precise device of the fork reactions and the common mechanism of DNA replication conserved among the prokaryotes and eukaryotes are examples of research themes that have long attracted investigators.4043). Several mechanisms have been proposed to explain how this might occur. Sadly, Reiji died of leukaemia in 1975, aged just 44. The discovery of Okazaki fragments has greatly advanced our Okazaki was born in Hiroshima, Japan. Louis 15 months (September 1960 to November 1961) and investigated novel nucleotide linked sugar compounds.69) In addition, there we learned micro-technology which could handle ul volume reagents and samples utilizing home-made glass micro pipettes. Immediately after I had lost Reiji, I received news about a serious challenge to the existence of Okazaki fragments. Thus, Rad59 promotes fork progression when Okazaki fragment processing is compromised and counteracts PCNA-K107 mediated cell cycle arrest.When we analyzed Okazaki fragments of prokaryotic cells based on this strategy, we were able to detect the 5-OH structure on some of Okazaki fragments. cdc9 rad59 double mutants did not alter PCNA ubiquitination but enhanced phosphorylation of the mediator of the replication checkpoint, Mrc1, indicative of increased replication fork stalling. To further understand how cells cope with nicks during replication, we utilized cdc9-1 in a genome-wide synthetic lethality screen and identified RAD59 as a strong negative interactor. Both enzymes reversed PCNA ubiquitination, arguing that the modification is likely triggered directly by nicks. To determine whether PCNA ubiquitination occurred in response to nicks or the lack of PCNA-DNA ligase interaction, we complemented cdc9 cells either with wild-type DNA ligase I or Chlorella virus ligase, the latter of which fails to interact with PCNA. In support of this notion, a pol30K107 mutation alleviated cell cycle arrest in cdc9 mutants. Most importantly, this signal is crucial to activate the S phase checkpoint, which promotes cell cycle arrest. The modification at K107 is catalyzed by the E2 variant Mms2 together with Ubc4 and the E3 ubiquitin ligase Rad5. cerevisiae as a model system, we uncovered a novel and conserved ubiquitination pathway that targets proliferating cell nuclear antigen (PCNA) at lysine 107 when DNA ligase I activity is inhibited. How cells monitor and suppress such accumulation of DNA damage that arises due to defective Okazaki fragment processing is unclear. 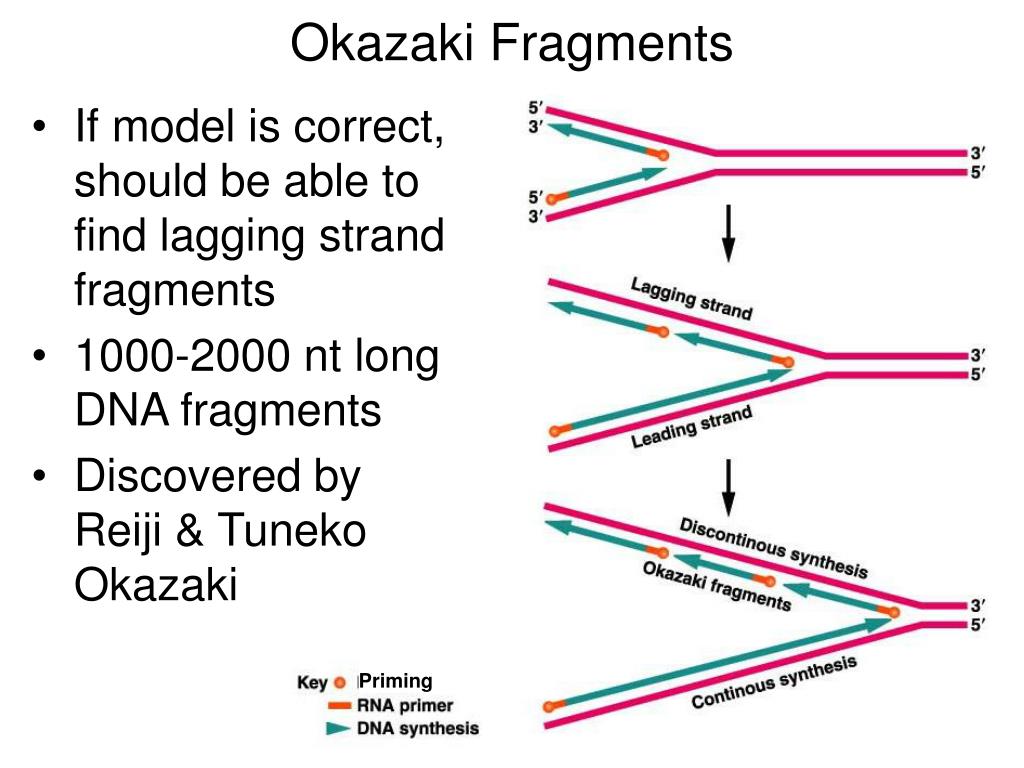
An individual harboring DNA ligase I mutations exhibited growth retardation, sunlight sensitivity, severe immunosuppression and developed lymphoma, indicating a link between defects in Okazaki fragment maturation and cancer predisposition. In humans, approximately 30 million Okazaki fragments are synthesized during every S phase and require further processing prior to DNA ligation. DNA ligase I, encoded by the CDC9 gene in Saccharomyces cerevisiae, is an essential enzyme that catalyzes the ligation of newly synthesized DNA on the lagging strand called Okazaki fragments. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
